 McGuffee SR, Smith DJ, Whitehouse I (2013) Quantitative, genome-wide analysis of eukaryotic replication initiation and termination. Smith DJ, Whitehouse I (2012) Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nasmyth KA (1977) Temperature-sensitive lethal mutants in the structural gene for DNA ligase in the yeast Schizosaccharomyces pombe. Johnston LH, Nasmyth KA (1978) Saccharomyces cerevisiae cell cycle mutant cdc9 is defective in DNA ligase. Distribution of functions between FEN1 AND DNA2. Nature 369:207–212Īyyagari R, Gomes XV, Gordenin DA et al (2003) Okazaki fragment maturation in yeast. Waga S, Stillman B (1994) Anatomy of a DNA replication fork revealed by reconstitution of SV40 DNA replication in vitro. Nature, 518:502–506.Nethanel T, Kaufmann G (1990) Two DNA polymerases may be required for synthesis of the lagging DNA strand of simian virus 40. Lagging-strand replication shapes the mutational landscape of the genome. 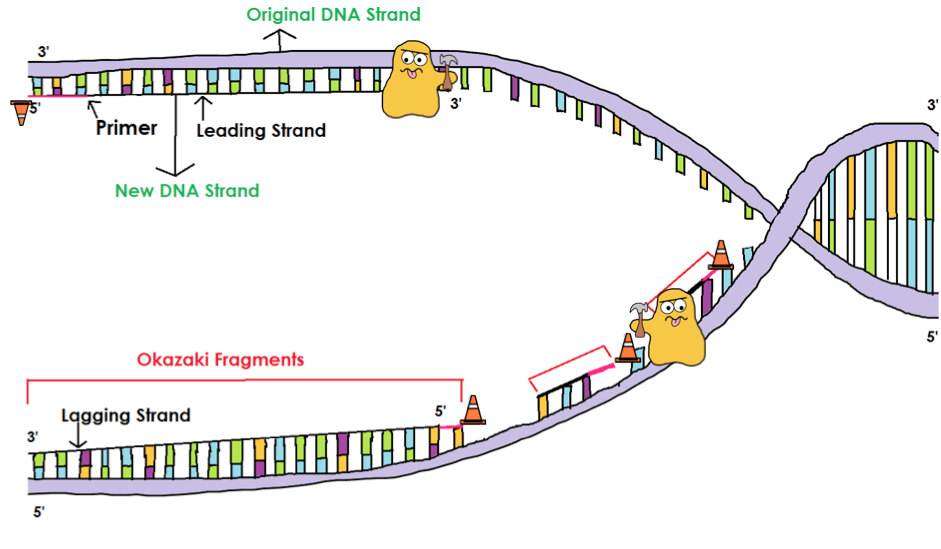
As a consequence these binding sites are biologically important mutational hotspots whose functional significance has been systematically underestimated by standard measures of sequence constraint. 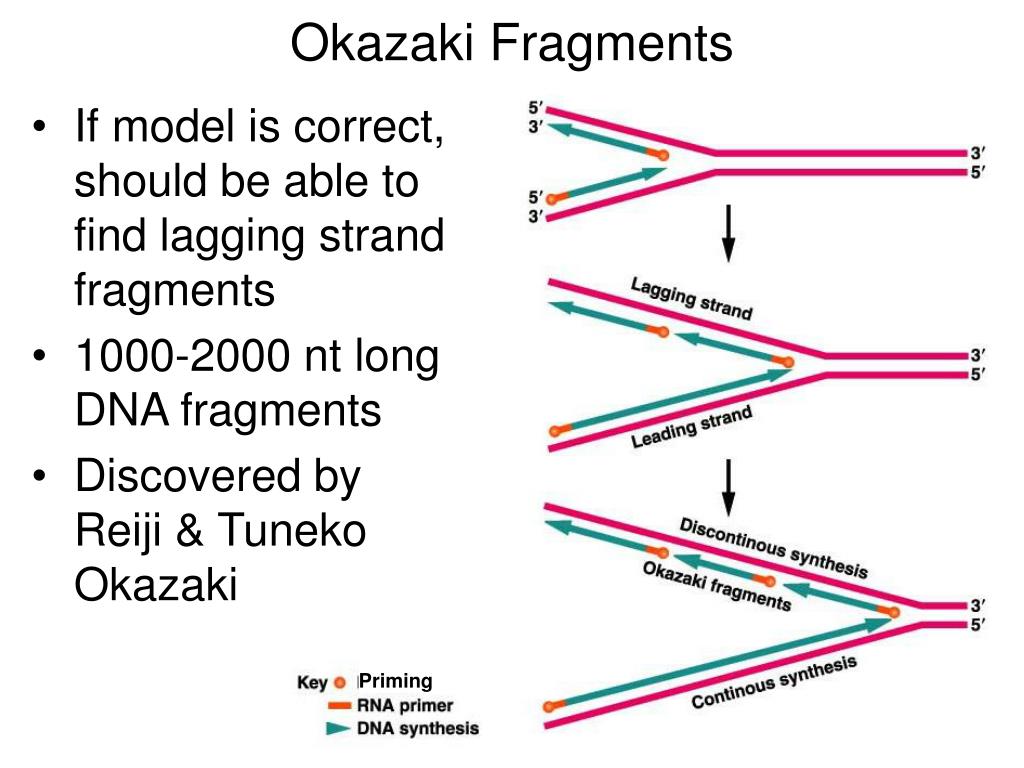
I propose that the rapid binding of some transcription factors to DNA following replication is required for nucleosome positioning or other important functions, however this incurs a cost in terms of locally elevated mutation rate adjacent to and within the sequence specific binding site. We observe that those footprints predicted as germline-specific manifest an elevated mutation signature. In the human genome I predict sites of preferential Pol α retention using DNase I hypersensitivity footprint data. I show that mutation hotspots adjacent to transcription factor binding sites are a conserved feature of eukaryotic genomes. Novel binding motifs are identified within the underlying DNA of these junctions that can be assigned to known strong and fast-binding transcription factors, previously implicated in the phasing of nucleosomes, such as Reb1. When these peaks are aligned and orientated, so that the direction of lagging strand replication is uniform, I find a peak in substitution rate immediately downstream of Okazaki junctions, precisely where Pol α tract retention is predicted to occur. I test this hypothesis using Okazaki fragment sequencing data from the yeast Saccharomyces cerevisiae to identify peaks in Okazaki junctions. 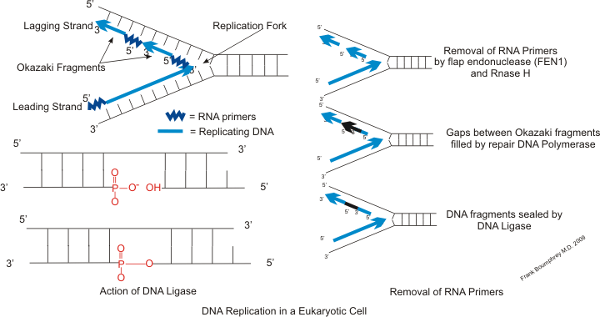
We present a mechanistic hypothesis to explain this locally elevated mutation rate and propose a role for lagging strand replication and its error-prone Pol α tract retention in the formation of these hotspots. I find that mutation rate is correlated genome wide with Okazaki junction frequency, suggesting that Okazaki junction processing may be error-prone. Some transcription factors are able to bind the DNA lagging strand during replication and act as a partial barrier to DNA polymerase, resulting in the accumulation of Okazaki fragment junctions adjacent to these sites. In eukaryotes both DNA strands are replicated simultaneously, the leading strand as a continuous stretch and the lagging strand as a series of discrete Okazaki fragments that are subsequently ligated together. In this work I investigate the relationship between DNA replication and apparent mutation hotspots adjacent to transcription factor binding sites. It is important to understand these patterns of mutation as they influence where deleterious mutations are likely to arise and how rapidly sequences are likely to accumulate change between species, a measure often used as a proxy for functional constraint. The rate of DNA mutation is known to fluctuate across the genome but the patterns of mutation rate variation and molecular causes are poorly defined. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
